Solvation and local motion of protein side chains studied by femto- and picosecond time-resolved fluorescence spectroscopy
Flexibility of biopolymers is one of the most characteristic properties of complex molecules, and it is intimately related to their functions. For proteins in aqueous environments, the relationship between the protein dynamics and that of the surrounding water molecules has attracted much interest. We applied time-resolved fluorescence spectroscopy to systematically quantify the relationship between solvation and local motion of protein side chains in the femto- to nanosecond time regime [1].
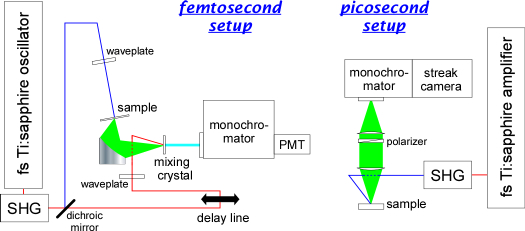
Femtosecond (left) and picosecond (right) time-resolved fluorescence
spectrometers used for dynamic Stokes shift experiments
The local motion of side chains was detected by monitoring the rotational relaxation of a fluorescent probe (acrylodan) attached to a protein (barstar), whereas the solvation dynamics were evaluated by measuring the dynamic Stokes shift of the fluorescent probe. The timescales of these motions varied depending on the location of the probe and the conformational state of the protein (native, unfolded, aggregated, and protofibril). However, in spite of the different absolute timescales of these dynamics (rotation: 0.1-3 ns, solvation: 20-300 ps), we observed a clear, positive correlation between them when different samples were compared. This result indicates strong coupling between the protein dynamics and the solvation dynamics.
This is a collaborative work with Prof. Krishnamoorthy and Prof. Udgaonkar (TIFR, India).
[1] “Exploration of the Correlation between Solvation Dynamics and Internal Dynamics of a Protein”, A. Jha, K. Ishii, J. B. Udgaonkar, T. Tahara and G. Krishnamoorthy, Biochemistry 50, 397-408 (2011).
